PDBcor: An automated correlation extraction calculator for multi-state protein structures - ScienceDirect
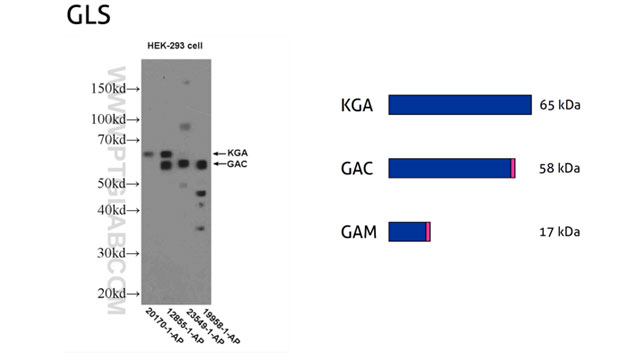
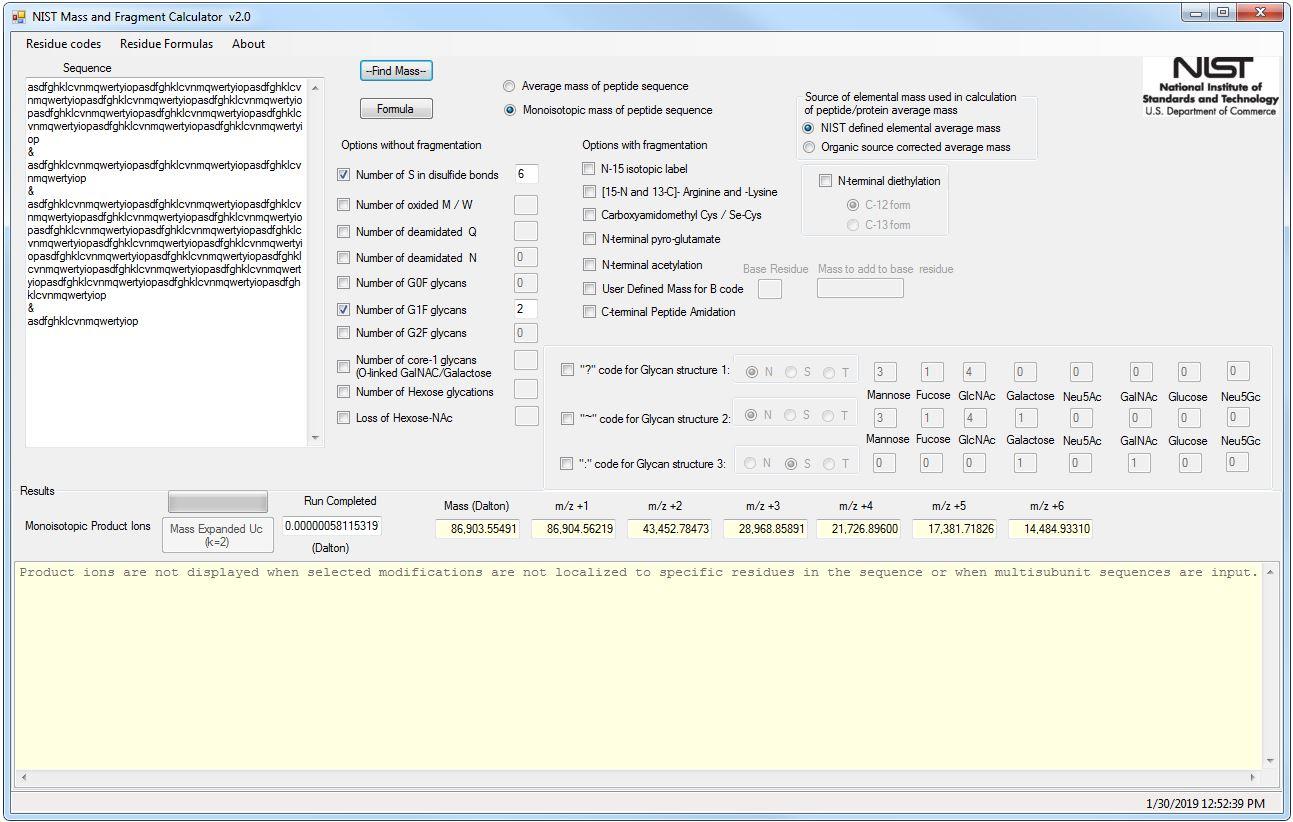
Why Does the Molecular Weight of My Protein Differ From the Theoretically Expected Weight? | Technology Networks
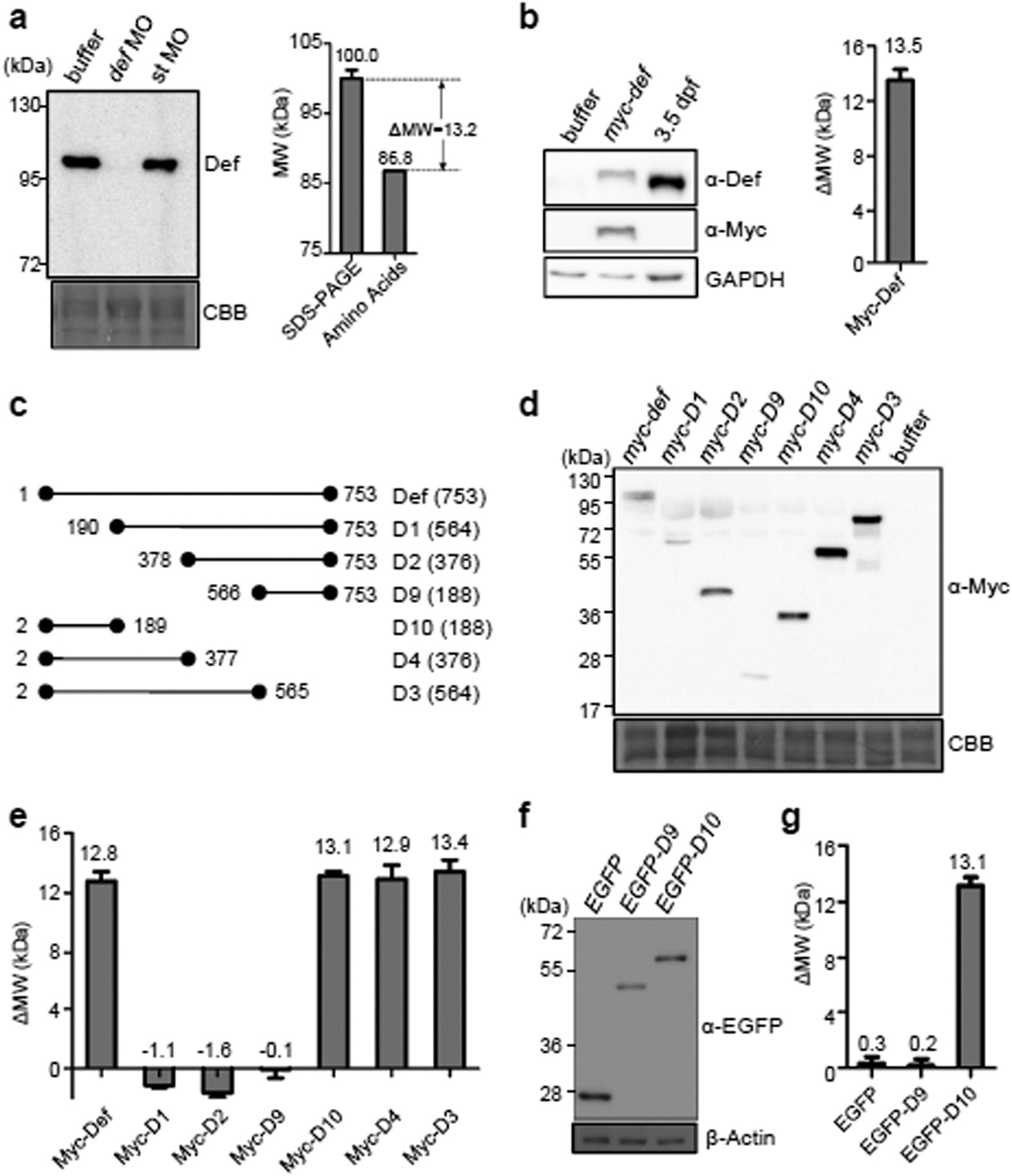
An equation to estimate the difference between theoretically predicted and SDS PAGE-displayed molecular weights for an acidic peptide | Scientific Reports
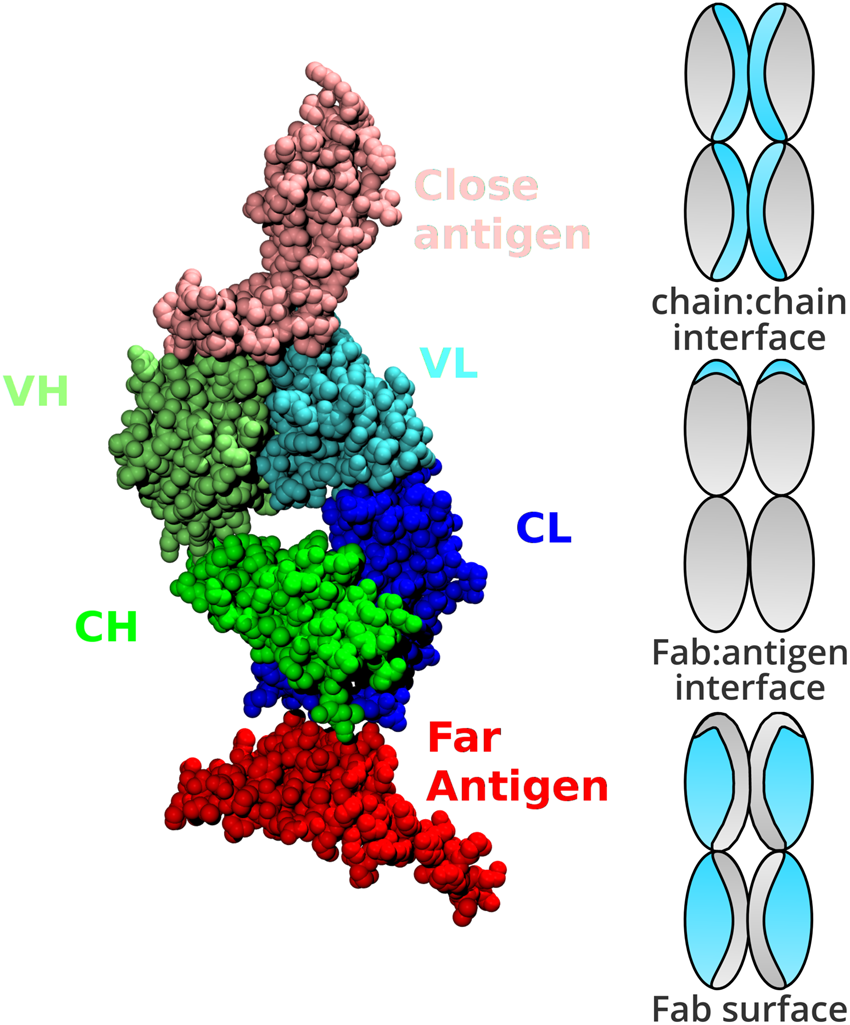
Web-based display of protein surface and pH-dependent properties for assessing the developability of biotherapeutics | Scientific Reports

SAXSMoW 2.0: Online calculator of the molecular weight of proteins in dilute solution from experimental SAXS data measured on a relative scale - Piiadov - 2019 - Protein Science - Wiley Online Library